I recently asked a friend who is a computational biologist at a major pharma company what her typical workday is like. By lunch, she had already toggled between 11 different applications: a gene database, two visualization tools, a statistical package, her electronic lab notebook, Slack, email, three browser tabs of papers, and a shared drive where her collaborator had uploaded the latest sequencing files. She spent 40 minutes reformatting a dataset so it would load into an analysis tool that didn’t speak the same file format as the instrument that generated it.
This is what passes for “doing life science” in 2025. The actual scientific thinking—forming hypotheses, interpreting results, designing the next experiment—gets squeezed into the gaps between tool-wrangling and waiting on specialists in a myriad of tools you don’t have time to master.
The bottleneck in modern biology isn’t ideas or even data. It’s human bandwidth. Labs generate terabytes weekly, but extracting insight requires stitching together fragmented software, translating between incompatible formats, and routing requests through scarce bioinformaticians who are perpetually backlogged. Researchers spend 80% of their time on what amounts to expensive glue work.
This is why I’m thrilled to announce that Menlo Ventures is co-leading Phylo’s $13.5 million seed round. Phylo is building what I’ve come to think of as the IDE for biology, a unified environment where scientists can finally work at the speed of their thinking.
A Structural Problem, Not a Software Problem
Software engineering faced a version of this problem decades ago. Code was written in one application, compiled in another, and debugged through painful printf statements after the fact. The tools existed, but they created friction at every handoff.
The integrated development environment changed everything, not by making any single tool better, but by collapsing the gaps between them. Write, run, debug, iterate, all in one place. Feedback loops that took hours compressed into seconds. The IDE didn’t replace programmers; it let them spend their time programming instead of managing tools.
Biology never got its equivalent. The reason isn’t lack of investment. Billions have poured into lab software. The problem is structural: A single research project might touch dozens of public databases, proprietary instruments, specialized analysis packages, and physical experiments at the bench. These systems were built by different teams, for different purposes, with different data models. No one designed them to interoperate, and no human could realistically master all of them.
But an AI agent trained on the documentation, file formats, and conventions of the entire biological software ecosystem? That’s a different story.
Intelligence as the Integration Layer
Phylo’s core insight is that foundation models can serve as the connective tissue biology has always lacked. The agent knows that Tool A’s output needs to be reshaped before Tool B will accept it. It understands the quirks of each database, the implicit conventions that experienced researchers accumulate over years. It holds context across a session the way a skilled collaborator would.
The technical foundation is a multi-agent architecture built for biological workflows, including planning, code generation, execution, and iterative refinement in a sandboxed environment. The platform already connects to hundreds of specialized tools, dozens of databases, and petabytes of biomedical data. Critically, every interaction generates structured traces that feed back into model improvement via reinforcement learning, creating a compounding data advantage.
The result is a workspace where a researcher can move from literature search to data analysis to experimental design without leaving the environment, without losing context, and without waiting in anyone’s queue. The scientist sets direction. The agents handle execution. The environment maintains continuity.
What Changes When the Friction Disappears
When I talk to researchers who’ve used Phylo, they describe something that sounds small but is actually profound: They can stay in the problem.
Today, the cognitive overhead of switching tools, reformatting data, and managing handoffs constantly interrupts scientific thinking. You have an idea, but before you can test it, you need to figure out which pipeline to use, debug why the script broke, and email someone who knows how to interpret the output. By the time you get the answer, you’ve lost the thread.
Phylo compresses that loop. A question that would have taken days of setup can be explored in an afternoon. The scientist’s role shifts from a technician manually executing each step to a director, focusing on what to investigate and how to interpret results.
The early traction suggests this resonates. In 72 days of research preview, Phylo attracted 8,400 users across 4,300 organizations, including 18 of the top 20 global pharma companies. Scientists at AstraZeneca, Novartis, and Astellas independently told us it’s among the most capable biology-specific agents they’ve evaluated. A pilot with another major pharma company compressed weeks-long analysis workflows into hours.
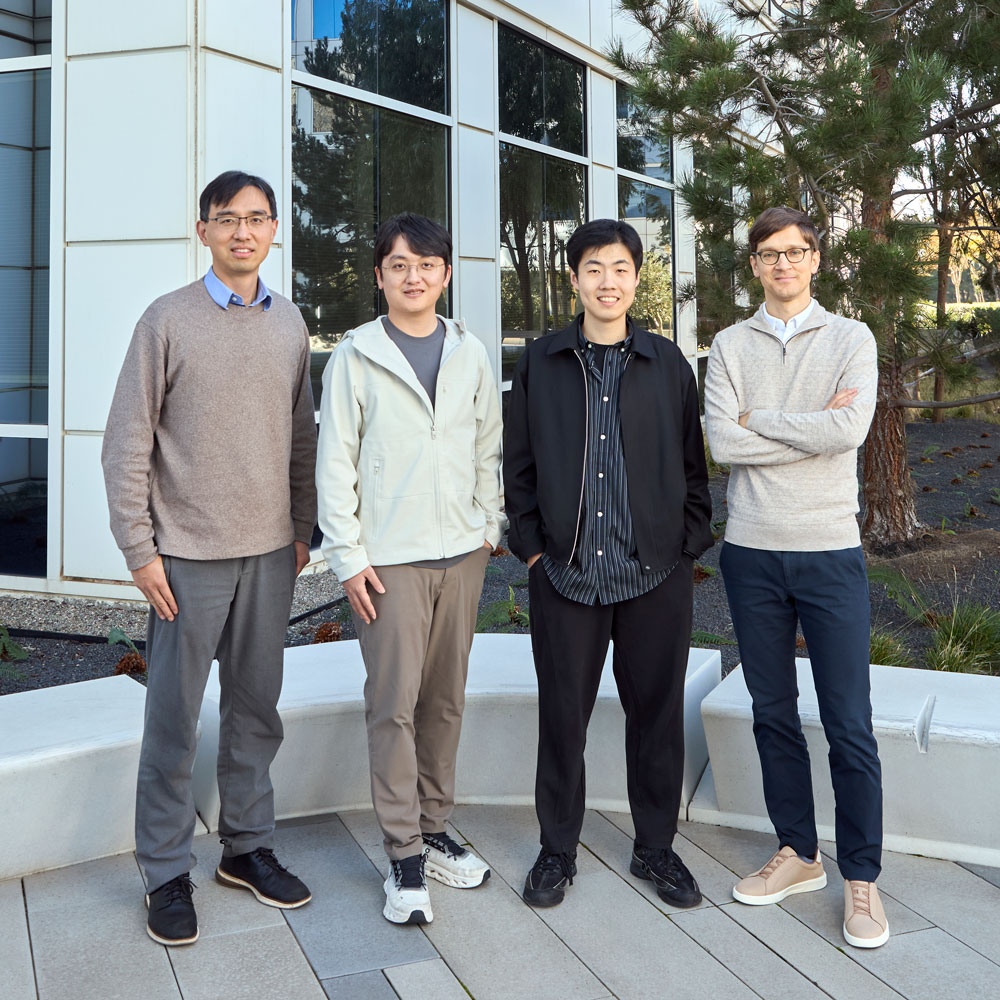
The Team

What convinced me to lead this round is the founders.
Kexin Huang recently completed his computer science Ph.D. at Stanford under Jure Leskovec, whose research group has shaped how the field thinks about graph neural networks and large-scale ML systems. Kexin’s own work focuses on AI for biomedical discovery, and he’s spent time in the trenches at GSK, Genentech, and Pfizer; he knows both the technical frontier and the messy reality of how pharma actually operates.
Yuanhao (Jerry) Qu recently completed his Ph.D. in genetics and pathology at Stanford, advised by Le Cong, who pioneered the use of CRISPR for gene editing. Jerry brings deep biological intuition and wet lab credibility that grounds the team’s technical ambitions in what scientists actually need.
I met them at a Stanford event last June before they even incorporated the company and have spent months since in regular working sessions, digging into the architecture, the roadmap, and the go-to-market. I even attended both of their Ph.D. defenses (unsurprisingly they both passed with flying colors!). They’re rigorous, intensely focused, and each individual has the rare combination of depth in both modern AI and modern biology, making them a perfect team to found this category-creating company.
Why Now
The timing of our investment isn’t accidental. Foundation models have reached a capability threshold where they can reason across heterogeneous tools and data formats in a way that wasn’t possible two years ago. Simultaneously, the cost of sequencing and other high-throughput methods has collapsed, generating more data than existing workflows can absorb. The mismatch between data abundance and analysis bandwidth has never been starker.
Phylo sits at that intersection: AI that’s finally capable enough, biology that’s finally data-rich enough, and researchers who are drowning in glue work and ready for something different.
Menlo Ventures is proud to co-lead this round. We’re betting that Phylo will do for biological research what the IDE did for software—not replace the scientists, but finally let them do science.
Matt Kraning is a partner at Menlo Ventures focused on AI, national defense, robotics, and cybersecurity. Before joining Menlo, he co-founded and served as CTO of Expanse, a category-creating data and cybersecurity company that Palo Alto Networks acquired for $1.25 billion. At Palo Alto Networks, he oversaw generative AI development…